Example: Use of bvector<> for k-mer fingerprint K should be short, no minimizers here (k < 24) More...
#include <assert.h>#include <stdlib.h>#include <math.h>#include <iostream>#include <vector>#include <list>#include <map>#include <algorithm>#include <utility>#include <memory>#include <future>#include <thread>#include <mutex>#include <atomic>#include "bm64.h"#include "bmalgo.h"#include "bmserial.h"#include "bmaggregator.h"#include "bmsparsevec_compr.h"#include "bmsparsevec_algo.h"#include "bmrandom.h"#include "bmdbg.h"#include "bmtimer.h"#include "bmundef.h"#include "dna_finger.h"#include "cmd_args.h"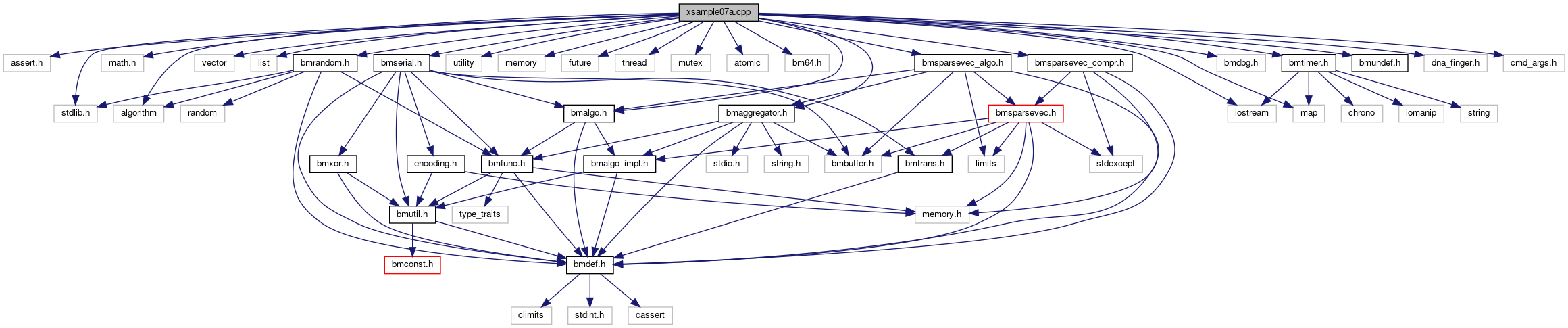
Go to the source code of this file.
Data Structures | |
| class | CSequenceColl |
| Collection of sequences and k-mer fingerprint vectors. More... | |
| class | CSeqGroup |
| Group (clustrer) of sequences. More... | |
| class | CSeqClusters |
| struct | CKMerAcc |
| Utility class to accumulate cahnges to cluster before commiting it (mutex syncronous operation) More... | |
Typedefs | |
| typedef bm::bvector | bvector_type |
| typedef std::vector< char > | vector_char_type |
| typedef bm::dynamic_heap_matrix< unsigned, bm::bvector<>::allocator_type > | distance_matrix_type |
| typedef std::vector< std::unique_ptr< bvector_type > > | bvector_ptr_vector_type |
| typedef bvector_type::size_type | bv_size_type |
| typedef std::vector< std::pair< bv_size_type, bv_size_type > > | bv_ranges_vector |
Functions | |
| template<typename FV > | |
| void | wait_for_slot (FV &futures, unsigned *parallel_cnt, unsigned concurrency) |
| wait for any opening in a list of futures used to schedule parallel tasks with CPU overbooking control More... | |
| static int | load_FASTA (const std::string &fname, CSequenceColl &seq_coll) |
| Load multi-sequence FASTA. More... | |
| static void | save_kmer_buffers (const std::string &fname, const CSequenceColl &seq_coll) |
| save k-mer vectors to a file More... | |
| static void | load_kmer_buffers (const std::string &fname, CSequenceColl &seq_coll) |
| Load k-mer vectors. More... | |
| bool | get_DNA_code (char bp, bm::id64_t &dna_code) |
| More... | |
| bool | get_kmer_code (const char *dna, size_t pos, unsigned k_size, bm::id64_t &k_mer) |
| Calculate k-mer as an unsigned long integer. More... | |
| char | int2DNA (unsigned code) |
| Translate integer code to DNA letter. More... | |
| void | translate_kmer (std::string &dna, bm::id64_t kmer_code, unsigned k_size) |
| Translate k-mer code into ATGC DNA string. More... | |
| template<typename BV > | |
| void | generate_k_mer_bvector (BV &bv, const vector_char_type &seq_vect, unsigned k_size, std::vector< bm::id64_t > &k_buf, const bm::id64_t chunk_size=400000000) |
| This function turns each k-mer into an integer number and encodes it in a bit-vector (presense vector) The natural limitation here is that integer has to be less tha 48-bits (limitations of bm::bvector<>) This method build a presense k-mer fingerprint vector which can be used for Jaccard distance comparison. More... | |
| std::atomic_ullong | k_mer_progress_count (0) |
| More... | |
| static void | generate_k_mers (CSequenceColl &seq_coll, unsigned k_size, size_t from, size_t to) |
| More... | |
| static void | generate_k_mers_parallel (CSequenceColl &seq_coll, unsigned k_size, unsigned concurrency) |
| More... | |
| void | resolve_duplicates (CSeqGroup &seq_group1, CSeqGroup &seq_group2, const CSequenceColl &seq_coll) |
| Resolve duplicate members between two groups. More... | |
| static void | compute_and_sim_row (unsigned *row, const bm::bvector<> *bv_i, size_t i, const std::vector< std::unique_ptr< bvector_type > > &k_mers_vect) |
| Compute similarity distances for one row/vector (1:N) of distance matrix. More... | |
| static void | compute_and_sim (distance_matrix_type &dm, const CSequenceColl &seq_coll, const bm::bvector<> &bv_mem, bm::bvector<>::size_type bv_mem_cnt, unsigned concurrency) |
| Compute similarity distances matrix (COUNT(AND(a, b)) More... | |
| static void | compute_seq_group_union (CSeqGroup &seq_group, const CSequenceColl &seq_coll) |
| Compute union (Universe) of all k-mers in the cluster group Implemented as a OR of all k-mer fingerprints. More... | |
| static void | compute_group (CSeqGroup &seq_group, const CSequenceColl &seq_coll, const bm::bvector<> &bv_exceptions, float similarity_cut_off) |
| More... | |
| static void | assign_to_best_cluster (CSeqClusters &cluster_groups, const CSequenceColl &seq_coll, const bm::bvector<> &bv_seq_ids, bm::bvector<>::size_type seq_id_from, bm::bvector<>::size_type seq_id_to) |
| Compute AND similarity to all available clusters assign to the most similar using cluster representative. More... | |
| static void | assign_to_best_cluster_union (CSeqClusters &cluster_groups, const CSequenceColl &seq_coll, const bm::bvector<> &bv_seq_ids, bm::bvector<>::size_type seq_id_from, bm::bvector<>::size_type seq_id_to) |
| Compute AND similarity to all available clusters assign to the most similar using UNION of k-mers in the cluster This is a more relaxed assignmnet, used when representative does not work. More... | |
| static void | compute_random_clusters (CSeqClusters &cluster_groups, const CSequenceColl &seq_coll, const bm::bvector<> &bv_total, bm::random_subset< bvector_type > &rsub, unsigned num_clust, float similarity_cut_off, unsigned concurrency) |
| Pick random sequences as cluster seed elements, try attach initial sequences based on weighted similarity measure. More... | |
| static void | compute_jaccard_clusters (CSeqClusters &seq_clusters, const CSequenceColl &seq_coll, unsigned num_clust, float similarity_cut_off, unsigned concurrency) |
| More... | |
| int | main (int argc, char *argv[]) |
| More... | |
Variables | |
| std::string | ifa_name |
| More... | |
| std::string | ikd_name |
| More... | |
| std::string | ikd_counts_name |
| std::string | kh_name |
| std::string | ikd_rep_name |
| std::string | ikd_freq_name |
| bool | is_diag = false |
| bool | is_timing = false |
| More... | |
| bool | is_bench = false |
| unsigned | ik_size = 8 |
| More... | |
| unsigned | parallel_jobs = 4 |
| More... | |
| unsigned | f_percent = 5 |
| bm::chrono_taker ::duration_map_type | timing_map |
| More... | |
Detailed Description
Example: Use of bvector<> for k-mer fingerprint K should be short, no minimizers here (k < 24)
Example loads FASTA file (large multi-molecule file is expected, builds a collection of k-mers for each molecule and runs clusterization algorithm on the input collection using set intersect (logical AND) as a similarity measure.
Example uses std::async for running parallel jobs.
Definition in file xsample07a.cpp.
Function Documentation
◆ assign_to_best_cluster()
|
static |
Compute AND similarity to all available clusters assign to the most similar using cluster representative.
- Examples
- xsample07a.cpp.
Definition at line 1245 of file xsample07a.cpp.
References CKMerAcc::add(), BM_DECLARE_TEMP_BLOCK, CKMerAcc::bv_kmers, CKMerAcc::bv_members, bm::count_and(), bm::deserialize(), CSequenceColl::get_buf(), CSeqClusters::get_group(), CSeqGroup::get_rep(), bm::bvector< Alloc >::enumerator::go_to(), CSeqClusters::groups_size(), CSeqGroup::merge_member_sync(), bm::bvector< Alloc >::set_allocator_pool(), and bm::bvector< Alloc >::iterator_base::valid().
Referenced by compute_jaccard_clusters().
◆ assign_to_best_cluster_union()
|
static |
Compute AND similarity to all available clusters assign to the most similar using UNION of k-mers in the cluster This is a more relaxed assignmnet, used when representative does not work.
- Examples
- xsample07a.cpp.
Definition at line 1315 of file xsample07a.cpp.
References CSeqGroup::add_member_sync(), BM_DECLARE_TEMP_BLOCK, CSeqGroup::count_and_union_sync(), bm::deserialize(), CSequenceColl::get_buf(), CSeqClusters::get_group(), bm::bvector< Alloc >::enumerator::go_to(), CSeqClusters::groups_size(), bm::bvector< Alloc >::set_allocator_pool(), and bm::bvector< Alloc >::iterator_base::valid().
Referenced by compute_jaccard_clusters().
◆ compute_and_sim()
|
static |
Compute similarity distances matrix (COUNT(AND(a, b))
- Examples
- xsample07a.cpp.
Definition at line 910 of file xsample07a.cpp.
References compute_and_sim_row(), CSequenceColl::deserialize_k_mers(), bm::random_subset< BV >::sample(), and wait_for_slot().
Referenced by CSeqClusters::elect_leaders().
◆ compute_and_sim_row()
|
static |
Compute similarity distances for one row/vector (1:N) of distance matrix.
- Examples
- xsample07a.cpp.
Definition at line 890 of file xsample07a.cpp.
References bm::bvector< Alloc >::count(), and bm::count_and().
Referenced by compute_and_sim().
◆ compute_group()
|
static |
- Examples
- xsample07a.cpp.
Definition at line 1102 of file xsample07a.cpp.
References CSeqGroup::add_member(), CSequenceColl::buf_size(), CSeqGroup::clear_member(), bm::bvector< Alloc >::count(), bm::deserialize(), bm::operation_deserializer< BV >::deserialize(), CSequenceColl::get_buf(), CSeqGroup::get_lead(), CSeqGroup::get_rep(), CSeqGroup::is_assigned(), bm::set_COUNT_AND, and bm::bvector< Alloc >::test().
Referenced by compute_random_clusters().
◆ compute_jaccard_clusters()
|
static |
- Examples
- xsample07a.cpp.
Definition at line 1420 of file xsample07a.cpp.
References assign_to_best_cluster(), assign_to_best_cluster_union(), CSequenceColl::buf_size(), CSeqClusters::clear_empty_groups(), CSeqClusters::compute_avg_count(), compute_random_clusters(), bm::bvector< Alloc >::count(), CSeqClusters::elect_leaders(), CSeqClusters::merge_from(), CSeqClusters::print_summary(), bm::rank_range_split(), CSeqClusters::resolve_duplicates(), bm::bvector< Alloc >::set_range(), CSeqClusters::take_group(), and CSeqClusters::union_all_groups().
Referenced by main().
◆ compute_random_clusters()
|
static |
Pick random sequences as cluster seed elements, try attach initial sequences based on weighted similarity measure.
- Examples
- xsample07a.cpp.
Definition at line 1374 of file xsample07a.cpp.
References CSeqClusters::add_group(), compute_group(), bm::bvector< Alloc >::first(), bm::random_subset< BV >::sample(), bm::bvector< Alloc >::iterator_base::valid(), and wait_for_slot().
Referenced by compute_jaccard_clusters().
◆ compute_seq_group_union()
|
static |
Compute union (Universe) of all k-mers in the cluster group Implemented as a OR of all k-mer fingerprints.
- Examples
- xsample07a.cpp.
Definition at line 973 of file xsample07a.cpp.
References BM_DECLARE_TEMP_BLOCK, bm::bvector< Alloc >::clear(), bm::deserialize(), CSequenceColl::get_buf(), CSeqGroup::get_kmer_union(), CSeqGroup::get_members(), bm::bvector< Alloc >::optimize(), and bm::bvector< Alloc >::iterator_base::valid().
Referenced by CSeqClusters::elect_leaders().
◆ generate_k_mer_bvector()
| void generate_k_mer_bvector | ( | BV & | bv, |
| const vector_char_type & | seq_vect, | ||
| unsigned | k_size, | ||
| std::vector< bm::id64_t > & | k_buf, | ||
| const bm::id64_t | chunk_size = 400000000 |
||
| ) |
This function turns each k-mer into an integer number and encodes it in a bit-vector (presense vector) The natural limitation here is that integer has to be less tha 48-bits (limitations of bm::bvector<>) This method build a presense k-mer fingerprint vector which can be used for Jaccard distance comparison.
- Parameters
-
bv - [out] - target bit-vector seq_vect - [out] DNA sequence vector k-size - dimention for k-mer generation k_buf - sort buffer for generated k-mers chunk_size - sort buffer size (number of k-mers per sort)
- Examples
- xsample07a.cpp.
Definition at line 474 of file xsample07a.cpp.
References bm::BM_SORTED, get_DNA_code(), and get_kmer_code().
Referenced by generate_k_mers().
◆ generate_k_mers()
|
static |
- Examples
- xsample07a.cpp.
Definition at line 561 of file xsample07a.cpp.
References BM_DECLARE_TEMP_BLOCK, generate_k_mer_bvector(), CSequenceColl::get_sequence(), k_mer_progress_count(), bm::bv_statistics::max_serialize_mem, bm::bvector< Alloc >::optimize(), bm::serializer< BV >::serialize(), bm::serializer< BV >::set_bookmarks(), CSequenceColl::set_buffer(), CSequenceColl::size(), and bm::bvector< Alloc >::size().
Referenced by generate_k_mers_parallel().
◆ generate_k_mers_parallel()
|
static |
- Examples
- xsample07a.cpp.
Definition at line 616 of file xsample07a.cpp.
References generate_k_mers(), k_mer_progress_count(), CSequenceColl::seq_size(), CSequenceColl::size(), and CSequenceColl::total_seq_size().
Referenced by main().
◆ get_DNA_code()
|
inline |
- Examples
- xsample07a.cpp.
Definition at line 374 of file xsample07a.cpp.
Referenced by generate_k_mer_bvector(), and get_kmer_code().
◆ get_kmer_code()
|
inline |
Calculate k-mer as an unsigned long integer.
- Returns
- true - if k-mer is "true" (not 'NNNNNN')
- Examples
- xsample07a.cpp.
Definition at line 402 of file xsample07a.cpp.
References get_DNA_code().
Referenced by generate_k_mer_bvector().
◆ int2DNA()
|
inline |
Translate integer code to DNA letter.
- Examples
- xsample07a.cpp.
Definition at line 429 of file xsample07a.cpp.
Referenced by translate_kmer().
◆ k_mer_progress_count()
| std::atomic_ullong k_mer_progress_count | ( | 0 | ) |
- Examples
- xsample07a.cpp.
Referenced by generate_k_mers(), and generate_k_mers_parallel().
◆ load_FASTA()
|
static |
Load multi-sequence FASTA.
- Examples
- xsample07a.cpp.
Definition at line 244 of file xsample07a.cpp.
References CSequenceColl::add_sequence().
Referenced by main().
◆ load_kmer_buffers()
|
static |
Load k-mer vectors.
- Examples
- xsample07a.cpp.
Definition at line 332 of file xsample07a.cpp.
References CSequenceColl::set_buffer().
Referenced by main().
◆ main()
| int main | ( | int | argc, |
| char * | argv[] | ||
| ) |
- Examples
- xsample07a.cpp.
Definition at line 1584 of file xsample07a.cpp.
References CSequenceColl::buf_size(), compute_jaccard_clusters(), generate_k_mers_parallel(), ifa_name, ik_size, ikd_name, is_timing, load_FASTA(), load_kmer_buffers(), parallel_jobs, parse_args(), bm::chrono_taker< TOut >::print_duration_map(), save_kmer_buffers(), CSequenceColl::size(), CSequenceColl::sync_buffers_size(), and timing_map.
◆ resolve_duplicates()
| void resolve_duplicates | ( | CSeqGroup & | seq_group1, |
| CSeqGroup & | seq_group2, | ||
| const CSequenceColl & | seq_coll | ||
| ) |
Resolve duplicate members between two groups.
- Examples
- xsample07a.cpp.
Definition at line 1155 of file xsample07a.cpp.
References bm::bvector< Alloc >::any(), bm::bvector< Alloc >::bit_and(), CSeqGroup::clear_member(), bm::operation_deserializer< BV >::deserialize(), CSequenceColl::get_buf(), CSeqGroup::get_lead(), CSeqGroup::get_members(), CSeqGroup::get_rep(), bm::set_COUNT_AND, and bm::bvector< Alloc >::iterator_base::valid().
◆ save_kmer_buffers()
|
static |
save k-mer vectors to a file
- Examples
- xsample07a.cpp.
Definition at line 294 of file xsample07a.cpp.
References CSequenceColl::get_buf(), CSequenceColl::get_buf_size(), and CSequenceColl::size().
Referenced by main().
◆ translate_kmer()
|
inline |
Translate k-mer code into ATGC DNA string.
- Parameters
-
dna - target string k_mer - k-mer code k_size -
- Examples
- xsample07a.cpp.
Definition at line 444 of file xsample07a.cpp.
References int2DNA().
◆ wait_for_slot()
| void wait_for_slot | ( | FV & | futures, |
| unsigned * | parallel_cnt, | ||
| unsigned | concurrency | ||
| ) |
wait for any opening in a list of futures used to schedule parallel tasks with CPU overbooking control
- Examples
- xsample07a.cpp.
Definition at line 111 of file xsample07a.cpp.
Referenced by compute_and_sim(), and compute_random_clusters().